FAIR Data Management#
The FAIR Guiding Principles for scientific data management and stewardship define how data and metadata should be structured to be Findable, Accessible, Interoperable, and Reusable. These principles rely on machine-actionable metadata, enabling data to be discovered, accessed, combined, and reused data with minimal human intervention.
To ensure unambiguous interpretation and reuse, metadata must be sufficiently detailed. In the Life Sciences, this is addressed through Minimum Information standards, which consist of two parts:
Community-defined reporting checklists for specific assays and data types
Standardized data formats for essential metadata to ensure machine-actionability
FAIR BY DESIGN#
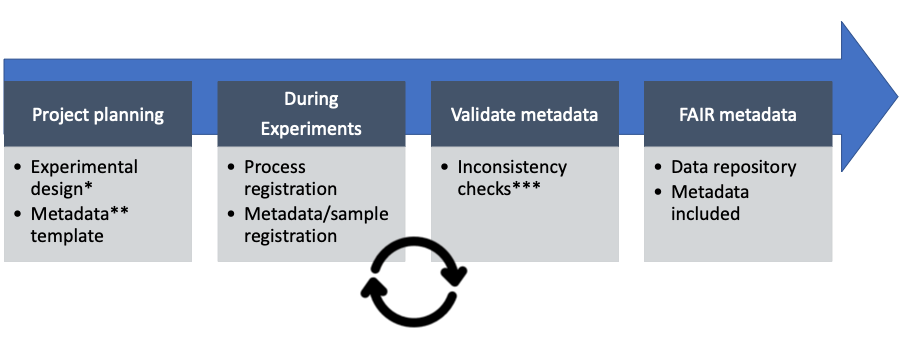
Working in a FAIR by Design manner ensures that projects are structured according to FAIR principles from their inception. These principles are applied throughout the project lifecyle to improve research quality.
This process is initiated by project planning and experimental design using the FAIR Data Station (https://fairds.fairbydesign.nl), where all relevant research metadata can be recorded. This recorded metadata is then transformed into a Linked Data file (RDF) which can be used for automated processing, data publishing and exploratory queries over a multitude of studies.
Role of ISA#
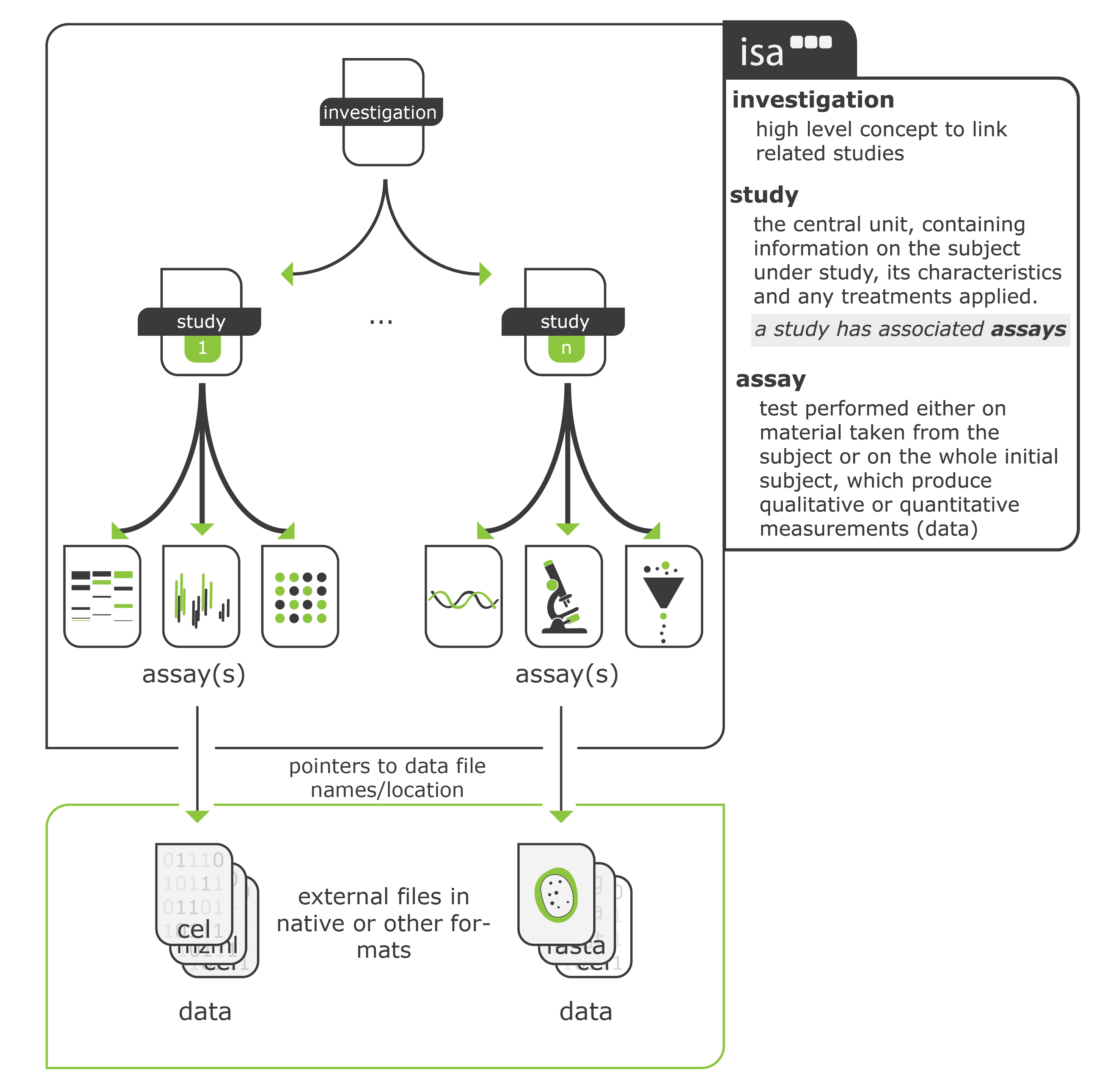
The core of our linked metadata system is based on the ISA format (http://isa-tools.org). ISA (Investigation, Study and Assay) is a metadata framework used to describe experiments in the life, environmental and biomedical sciences. It consists of three core conepts:
Investigation: the overall project context
Study: a unit of research within the project
Assay: analytical measurements performed within a study These concepts are incorporated and extended with additional standards such as JERM, MIAPPE and MIxS.
Ontologies#
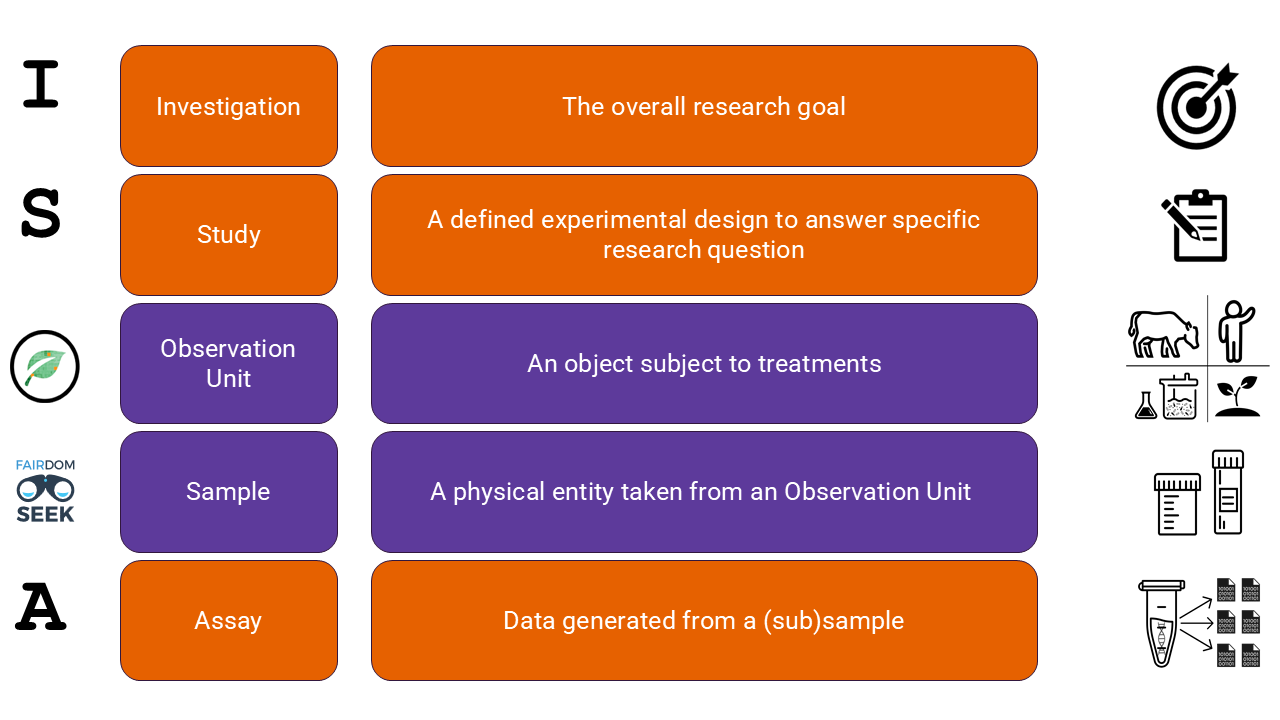
A set of ontologies is used to support the metadata model. This model is based on the ISA structure, where URIs from ISA are mapped to the Just Enough Results Model. Through JERM, a Sample class was added to improve the alignment with the FAIRDOM Hub and to enable grouping of multiple investigations under one funding agency (the project).
In biological studies, samples play a central role, and multiple subjects or environments are oftentimes analyised within a single study. The addition of Observation Unit from the MIAPPE ontology resolves the missing association between sample and study.
API#
The ontology is defined using OWL and ShEx, enabling Empusa to generate a Java API for integration purposes. The API (http://download.systemsbiology.nl/#/unlock/) is used to create, manage and query RDF datasets. Within the FAIR Data Station (https://fairds.fairbydesign.nl) the ontology is used to validate metadata and generate structured RDF data from Excel-based metadata sheets.
The metadata information of these RDF datasets is subsequently used to drive the computational work, leveraging the Common Workflow Language (CWL) to process raw data to structured results.
